pyNastran/op2/op2¶
This is the pyNastran.op2.op2.rst file.
op2 Module¶
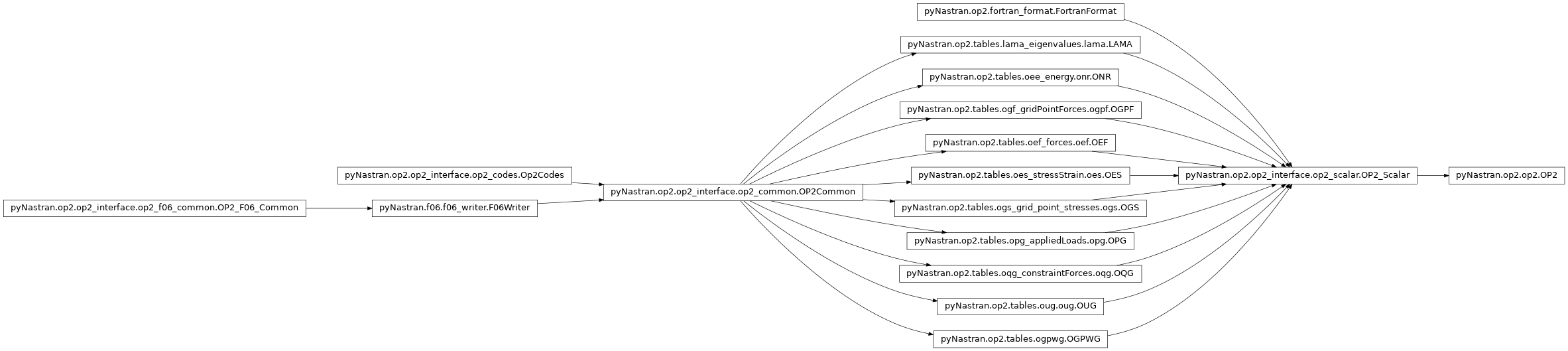
Defines the main OP2 class. Defines:
- read_op2(op2_filename=None, combine=True, subcases=None,
exclude_results=None, include_results=None, log=None, debug=True, debug_file=None, build_dataframe=None, skip_undefined_matrices=True, mode=’msc’, encoding=None)
OP2(debug=True, log=None, debug_file=None, mode=’msc’) - build_dataframe() - combine_results(combine=True) - create_objects_from_matrices() - object_attributes(mode=’public’, keys_to_skip=None, filter_properties=False) - object_methods(mode=’public’, keys_to_skip=None) - print_subcase_key() - read_op2(op2_filename=None, combine=True, build_dataframe=None,
skip_undefined_matrices=False, encoding=None)
- set_mode(mode)
- transform_displacements_to_global(i_transform, coords, xyz_cid0=None, debug=False)
- transform_gpforce_to_global(nids_all, nids_transform, i_transform, coords, xyz_cid0=None)
-
class
pyNastran.op2.op2.OP2(debug=True, log=None, debug_file=None, mode=None)[source]¶ Bases:
pyNastran.op2.op2_interface.op2_scalar.OP2_ScalarInitializes the OP2 object
Parameters: - debug : bool; default=False
enables the debug log and sets the debug in the logger
- log : Log()
a logging object to write debug messages to
(.. seealso:: import logging)
- debug_file : str; default=None (No debug)
sets the filename that will be written to
- mode : str; default=None -> ‘msc’
{msc, nx}
-
assert_op2_equal(self, op2_model, skip_results=None, stop_on_failure=True, debug=False)[source]¶ Diffs the current op2 model vs. another op2 model.
Parameters: - op2_model : OP2()
the model to compare to
- skip_results : List[str]; default=None -> []
results that shouldn’t be compred
- stop_on_failure : bool; default=True
True : Crashes if they’re not equal False : Go to the next object
- debug : bool; default=False
give slightly more debugging messages
Returns: - is_equal : bool
are the objects equal?
Raises: - AssertionError/ValueError : stop_on_failure=True and and error occurred
- NotImplementedError : this is a sign of an unsupported object
-
build_dataframe(self)[source]¶ Converts the OP2 objects into pandas DataFrames
Todo
fix issues with: - RealDisplacementArray - RealPlateStressArray (???) - RealPlateStrainArray (???) - RealCompositePlateStrainArray (???)
-
combine_results(self, combine=True)[source]¶ we want the data to be in the same format and grouped by subcase, so we take
stress = { # isubcase, analysis_code, sort_method, count, superelement_adaptivity_index, pval_step (1, 2, 1, 0, 'SUPERELEMENT 0', '') : result1, (1, 2, 1, 0, 'SUPERELEMENT 10', '') : result2, (1, 2, 1, 0, 'SUPERELEMENT 20', '') : result3, (2, 2, 1, 0, 'SUPERELEMENT 0', '') : result4,
- code = (isubcase, analysis_code, sort_method, count, ogs,
- superelement_adaptivity_index, pval_step)
}
and convert it to:
stress = { 1 : result1 + result2 + results3, 2 : result4, }
-
create_objects_from_matrices(self)[source]¶ - creates the following objects:
- monitor3 : MONPNT3 object from the MP3F matrix
- monitor1 : MONPNT1 object from the PMRF, PERF, PFRF, AGRF, PGRF, AFRF matrices
-
export_hdf5_file(self, hdf5_file, exporter=None)[source]¶ Converts the OP2 objects into hdf5 object
Parameters: - hdf5_file : H5File()
an h5py object
- exporter : HDF5Exporter; default=None
unused
- TODO: doesn’t support:
- BucklingEigenvalues
-
export_hdf5_filename(self, hdf5_filename)[source]¶ Converts the OP2 objects into hdf5 object
- TODO: doesn’t support:
- BucklingEigenvalues
-
include_exclude_results(self, exclude_results=None, include_results=None)[source]¶ Sets results to include/exclude
Parameters: - exclude_results / include_results : List[str] / str; default=None
a list of result types to exclude/include one of these must be None
-
is_geometry¶
-
load_hdf5_file(self, h5_file, combine=True)[source]¶ Loads an h5 file object into an OP2 object
Parameters: - h5_file : H5File()
an h5py file object
- combine : bool; default=True
runs the combine routine
-
load_hdf5_filename(self, hdf5_filename, combine=True)[source]¶ Loads an h5 file into an OP2 object
Parameters: - hdf5_filename : str
the path to the an hdf5 file
- combine : bool; default=True
runs the combine routine
-
object_attributes(self, mode='public', keys_to_skip=None, filter_properties=False)[source]¶ List the names of attributes of a class as strings. Returns public attributes as default.
Parameters: - mode : str
defines what kind of attributes will be listed * ‘public’ - names that do not begin with underscore * ‘private’ - names that begin with single underscore * ‘both’ - private and public * ‘all’ - all attributes that are defined for the object
- keys_to_skip : List[str]; default=None -> []
names to not consider to avoid deprecation warnings
Returns: - attribute_names : List[str]
sorted list of the names of attributes of a given type or None if the mode is wrong
-
object_methods(self, mode='public', keys_to_skip=None)[source]¶ List the names of methods of a class as strings. Returns public methods as default.
Parameters: - obj : instance
the object for checking
- mode : str
defines what kind of methods will be listed * “public” - names that do not begin with underscore * “private” - names that begin with single underscore * “both” - private and public * “all” - all methods that are defined for the object
- keys_to_skip : List[str]; default=None -> []
names to not consider to avoid deprecation warnings
Returns: - method : List[str]
sorted list of the names of methods of a given type or None if the mode is wrong
-
read_op2(self, op2_filename=None, combine=True, build_dataframe=None, skip_undefined_matrices=False, encoding=None)[source]¶ Starts the OP2 file reading
Parameters: - op2_filename : str (default=None -> popup)
the op2_filename
- combine : bool; default=True
True : objects are isubcase based False : objects are (isubcase, subtitle) based;
will be used for superelements regardless of the option
- #load_as_h5 : default=False
#loads the op2 as an h5 file to save memory #stores the result.element/data attributes in h5 format
- build_dataframe : bool (default=None -> True if in iPython, False otherwise)
builds a pandas DataFrame for op2 objects
- skip_undefined_matrices : bool; default=False
True : prevents matrix reading crashes
- encoding : str
the unicode encoding (default=None; system default)
-
transform_displacements_to_global(self, icd_transform, coords, xyz_cid0=None, debug=False)[source]¶ Transforms the
dataof displacement-like results into the global coordinate system for those nodes with different output coordinate systems. Takes indicies and transformation matricies for nodes with their output in coordinate systems other than the global.Used in combination with
BDF.get_displacement_indexParameters: - icd_transform : dict{int cid
Dictionary from coordinate id to index of the nodes in
BDF.point_idsthat their output (CD) in that coordinate system.- coords : dict{int cid :Coord()}
Dictionary of coordinate id to the coordinate object Use this if CD is only rectangular Use this if CD is not rectangular
- xyz_cid0 : (nnodes+nspoints, 3) float ndarray
the nodes in the global frame Don’t use this if CD is only rectangular Use this if CD is not rectangular
- debug : bool; default=False
developer debug
- .. warning:: only works if all nodes are included…
test_pynastrangui isat_tran.dat isat_tran.op2 -f nastran- .. note:: Nastran has this concept of a basic (cid=0) and global (cid=cd)
coordinate system. They occur at the same time. Basic is for positions/properties, while global is for result outputs.
- pyNastran’s OP2 interface uses:
- cd=0 for global frames
- cd>0 are local frames
- pyNastran’s BDF interface uses:
- cp=0 for global frames
- cp>0 are local frames
-
transform_gpforce_to_global(self, nids_all, nids_transform, icd_transform, coords, xyz_cid0=None)[source]¶ Transforms the
dataof GPFORCE results into the global coordinate system for those nodes with different output coordinate systems. Takes indicies and transformation matricies for nodes with their output in coordinate systems other than the global.Used in combination with
BDF.get_displacement_indexParameters: - nids_all : ???
???
- nids_transform : dict{int cid
Dictionary from coordinate id to corresponding node ids.
- icd_transform : dict{int cid
Dictionary from coordinate id to index of the nodes in
BDF.point_idsthat their output (CD) in that coordinate system.- coords : dict{int cid :Coord()}
Dictionary of coordinate id to the coordinate object Use this if CD is only rectangular Use this if CD is not rectangular
- xyz_cid0 : ???
required for cylindrical/spherical coordinate systems
-
pyNastran.op2.op2.read_op2(op2_filename=None, combine=True, subcases=None, exclude_results=None, include_results=None, log=None, debug=True, debug_file=None, build_dataframe=None, skip_undefined_matrices=True, mode=None, encoding=None)[source]¶ Creates the OP2 object without calling the OP2 class.
Parameters: - op2_filename : str (default=None -> popup)
the op2_filename
- combine : bool; default=True
True : objects are isubcase based False : objects are (isubcase, subtitle) based;
will be used for superelements regardless of the option
- subcases : List[int, …] / int; default=None->all subcases
list of [subcase1_ID,subcase2_ID]
- exclude_results / include_results : List[str] / str; default=None
a list of result types to exclude/include one of these must be None
- build_dataframe : bool (default=None -> True if in iPython, False otherwise)
builds a pandas DataFrame for op2 objects
- skip_undefined_matrices : bool; default=False
True : prevents matrix reading crashes
- debug : bool; default=False
enables the debug log and sets the debug in the logger
- log : Log()
a logging object to write debug messages to (.. seealso:: import logging)
- mode : str; default=None -> ‘msc’
the version of the Nastran you’re using {nx, msc, optistruct}
- debug_file : str; default=None (No debug)
sets the filename that will be written to
- encoding : str
the unicode encoding (default=None; system default)
Returns: - model : OP2()
an OP2 object
Todo
creates the OP2 object without all the read methods ..
Note
this method will change in order to return an object that does not have so many methods